Using Modules in R
When a code base grows we may think of using several files first and then source them. Functions, of course, are rightfully advocated to new R users, and are the essential building block. Packages are then, already, the next level of abstraction we have to offer. With the modules package I want to provide something in between: local namespace definitions without, or within R packages. We find this feature implemented in various ways and languages: classes, namespaces, functions, packages, and sometimes also modules. Python, Julia, F#, Scala, and Erlang (and more) are languages which use modules as a language construct. Some of them in parallel to classes and packages.
More Than a Function, Less Than a Package
Just like classes and objects, modules present a way to group functions into one entity. They behave as a first class citizen in the sense that they can be treated like any other data structure in R:
- they can be created anywhere, including inside another module,
- they can be passed to functions,
- and returned from functions.
In addition they provide:
- local namespace features by declaring imports and exports
- encapsulation by introducing a local scope,
- code reuse by various modes of composition,
- interchangeability with other modules implementing the same interface.
The most important part for me is the freedom to create a local scope, or context, in which I can safely reuse verbs as function names. This is so important to me because naming things is so terribly difficult; many verbs have a different meaning in a different context; and a lot of expressive verbs are already used by base R or the tidyverse and I want to avoid name clashes. When you are working inside a package or in the REPL, aka console, R only knows one context: all values are bound to names in the global environment (or package scope). Modules present a way to create a new context and reuse names, among some other useful features. To make one thing clear: modules do not aim at being a substitute for packages. Instead we may use them to create local namespaces inside or outside of a package. On a scale of abstraction they are located between functions and packages: modules can comprise functions, sometimes data. Packages can comprise functions, data, and modules (furthermore documentation, tests, and are a solid way to share code).
A Simple Module Definition
Let’s look at an example:
# install.packages("modules")
# vignette("modulesInR", "modules")
graphics <- modules::module({
modules::import("ggplot2")
barplot <- function(df) {
## my example barplot
## df (data.frame) a data frame with group and count columns
ggplot(df, aes(group, count)) +
geom_bar(stat = "identity") +
labs(x = "Some group variable", y = "Absolute frequency")
}
})
df <- data.frame(
# Just some sample data
group = sample(LETTERS, 5),
count = round(runif(5, max = 100))
)
graphics$barplot(df)
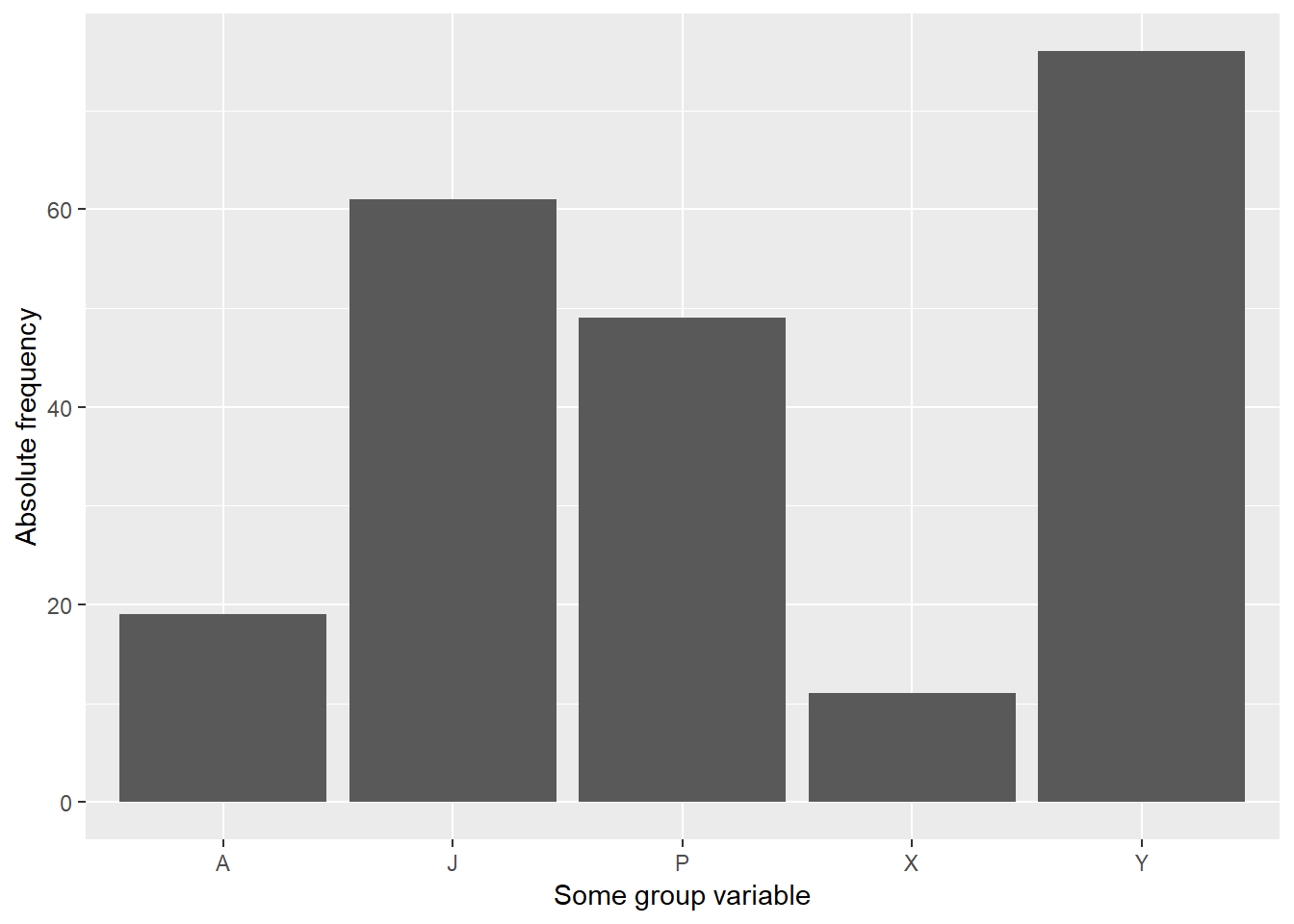
First we instantiate a new module using modules::module(). What you should recognize is that you can simply put your function definitions into it; here we define the function barplot. We then can call that function by qualifying its name with graphics$barplot. The graphics module is just a list and as such can be passed around:
class(graphics)
## [1] "module" "list"
graphics
## barplot:
## function(df)
## ## my example barplot
## ## df (data.frame) a data frame with group and count columns
Something which is important about modules is that they aim to create a local scope, or a mini namespace. As such they support to define imports and exports. So far we have made all names exported by ggplot2 available to the scope of the module. We may gain finer control of the names available to the module as illustrated below:
graphics <- modules::module({
modules::import("ggplot2", "aes", "geom_bar", "ggplot", "labs")
modules::export("barplot")
barplot <- function(df) {
## my example barplot
## df (data.frame) a data frame with a group and count column
ggplot(df, aes(group, count)) +
geom_bar(stat = "identity") +
labs(x = "Some group variable", y = "Absolute frequency")
}
})
We do not have to use the modules::module function to create a module. When we write scripts it might be more convenient, to instead spread our codebase accross files. Each file can then act as a module. If you compare this to source the advantage is, that each file can have its own set of imports while avoiding any name clashes between them. Furthermore modules only export what is necessary and we avoid to spam the global environment with intermediate objects and function definitions:
graphics.R
modules::import("ggplot2", "aes", "geom_bar", "ggplot", "labs")
modules::export("barplot")
barplot <- function(df) {
## my example barplot
## df (data.frame) a data frame with a group and count column
ggplot(df, aes(group, count)) +
geom_bar(stat = "identity") +
labs(x = "Some group variable", y = "Absolute frequency")
}
Then we load this module using:
graphics <- modules::use("graphics.R")
graphics$barplot
When Can Modules be Useful?
Why would you want this? All too often we end up spending much more time than we should maintaining badly written code bases. And while the solution often is: "write a package!", "use a style guide!", "structure!" – we just tend to respond "We can pretify later, I just need to finish this up first." Modules aim to provide a lightweight solution to a couple of problems I often see in code bases:
Imagine you write a report involving some sort of data analysis. Of course you use knitr or Sweave. So you combine your writing and coding in one file. After a while the report grows beyond the length of a blog post and is too long for one file. So as a faithful useR you start to write functions. Maybe now you also start to have several R scripts or multiple Rmd files where you put your function definitions. Of course your library statements go into some sort of header block, right? Your functions depending on these library statements are separated into files or can be found possibly thousands of lines down the road. After some time, maybe month, and the effort of several authors, you discover that your report only compiles in a fresh R session. Never twice in a row, you get different results then. Someone decided to add library statements into the source files. When you remove them and place them on top in that header section you are supposed to have, something breaks. When you remove some variables you do not need anymore something down the road breaks. You do not really know why and you also promised that you would get an update of the report ready: this afternoon. So you better leave everything unchanged and just get it done, somehow.
In those kind of situations we missed the point to get the codebase organized. It is uncontrolled growth and happens when we do not pay attention. It happens more often than we like to admit, and it is also avoidable. Aside from all good practices you can and should apply, modules offer:
- A safe(r) way to
sourcefiles in one R session - Avoid name clashes between packages: (a) by importing single functions and (b) by only importing them locally
- Avoid name clashes between different parts (maybe files) of your codebase. In many cases you rely on objects in your global workspace. With modules you can make them local
- Provide interfaces and hide the implementation
A Tool to Create Boundaries
Before we end up in those situation, what can we do? We need to create boundaries. We need to be explicit which dependencies are needed and where they are needed. We need also to be explicit which objects are allowed to be used in later parts of the analysis. When we manage to create boundaries between the parts of our programs, or between the parts of a report, we reduce the scope we need to understand when writing, maintaining - aka debugging - or extending any of it. We also take control of how much coupling between the parts we allow. The list goes on: boundaries are a good thing, in programming.
There are a number of tools and a variety of concepts you can use to create boundaries. Modules can be one tool in your bag. They allow the use of local imports: ggplot2 in the above code was never attached to the search path of the main R session. They allow us to expose only what is necessary and thus help us protect private objects. Modules can sit in their own files and folders and can be loaded, without changing anything in the global environment. Modules are containers where we can put things that are related. They can help us to keep things clean and impose structure. Sometimes I feel that writing functions for a package is like putting all the things I have into a storage room. Using modules, I additionally have boxes.
Final Remarks
Writing all our code in some class definition does not mean we do object orientation. Writing a lot of functions does not mean we do functional programming. Just by putting all your code into a module will not solve anything. Only when we know and understand the problems we try to solve can we pick the right tool. It is just plain hard to know in advance what the right structure is for your problem. Also it is not always obvious when the small script should have had a package to back it up. But we have to improve on this! Impose structure, decouple components, follow best practices and get organized! For these things, I hope that modules can be of use. Feel free to open an issue on GitHub if you have any questions.